Extract parameter simulations from the joint precision matrix
Source:R/gather-spread.R
gather_sims.Rdspread_sims() returns a wide-format data frame. gather_sims() returns a
long-format data frame. The format matches the format in the tidybayes
spread_draws() and gather_draws() functions.
Arguments
- object
Output from
sdmTMB().- nsim
The number of simulation draws.
Value
A data frame. gather_sims() returns a long-format data frame:
.iteration: the sample ID.variable: the parameter name.value: the parameter sample value
spread_sims() returns a wide-format data frame:
.iteration: the sample IDcolumns for each parameter with a sample per row
Examples
m <- sdmTMB(density ~ depth_scaled,
data = pcod_2011, mesh = pcod_mesh_2011, family = tweedie())
head(spread_sims(m, nsim = 10))
#> .iteration X.Intercept. depth_scaled range phi tweedie_p sigma_O
#> 1 1 2.912963 -0.4786992 64.038443 14.64030 1.608921 1.813936
#> 2 2 2.940852 -0.5085643 53.287196 15.10246 1.584533 1.740709
#> 3 3 2.368351 -0.6906364 20.392355 14.00913 1.569154 2.769780
#> 4 4 2.784025 -0.5624740 9.914927 15.61474 1.582818 3.090263
#> 5 5 2.863775 -0.5143786 27.041172 15.34518 1.585760 3.093743
#> 6 6 2.950199 -0.5919163 38.234861 15.99667 1.575896 2.002849
head(gather_sims(m, nsim = 10))
#> .iteration .variable .value
#> 1 1 X.Intercept. 3.273034
#> 2 2 X.Intercept. 2.980942
#> 3 3 X.Intercept. 2.570352
#> 4 4 X.Intercept. 2.934931
#> 5 5 X.Intercept. 3.093088
#> 6 6 X.Intercept. 2.769836
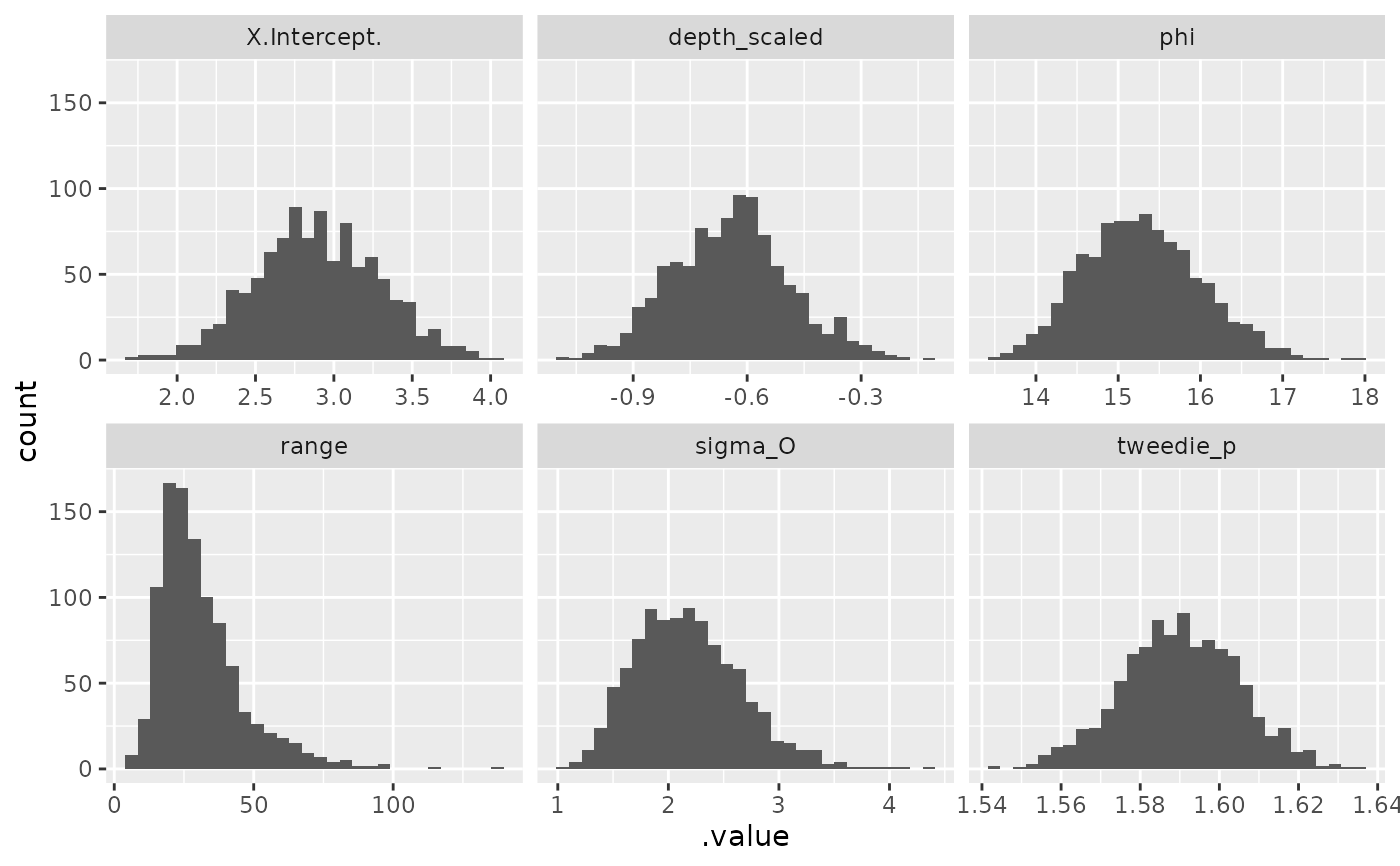
samps <- gather_sims(m, nsim = 1000)
if (require("ggplot2", quietly = TRUE)) {
ggplot(samps, aes(.value)) + geom_histogram() +
facet_wrap(~.variable, scales = "free_x")
}
#> `stat_bin()` using `bins = 30`. Pick better value `binwidth`.